
CONIFIR Project
Genetic origin and structural setting of Douglas-fir planted forests in Italy for their management, conservation and valorization
MUR-PRIN 2022 Project - Prot. 20225WRLS3

MUR-PRIN 2022 Project - Prot. 20225WRLS3

Rendering of an Italian Douglas-fir stand (Monti Cimini, VT, Italy) based on ALS scan (courtesy of: N. Puletti, CREA-FL, Italy).
The introduction of non-native tree species in Europe have led to the establishment of extensive forest stands which are currently “naturalised”, thus becoming part of the national forest heritage. However, this process could trigger a “genetic bottleneck” in the introduced populations, involving a loss of genetic variation and evolutionary potential needed to adapt to environmental changes (Sakai et al. 2001). To overcome that, non-native tree species have been often introduced (or re-introduced) from genetically differentiated source populations, thereby ensuring suitable genetic diversity and adaptability of the introduced plant material (Rijal et al. 2015).
Douglas-fir [Pseudotsuga menziesii (Mirb.) Franco] is a long-lived tree species native to the western coast of North America and Rocky Mountains and it is probably the most relevant example of non-native forest tree species introduced in Europe since the 1850s (Schmid et al. 2014). The huge native range of Douglas-fir can be divided into two geographically distinct varieties which differ morphologically and ecologically: P. menziesii var. menziesii (coastal variety), and P. menziesii var. glauca (Rocky Mountain - interior variety). Furthermore, within the var. glauca a third genetic lineage (Mexican lineage) was identified by sequencing chloroplast DNA (Wei et al. 2011). This species has been introduced multiple times in Europe from different geographic areas of its native range, and has been widely planted across different environments in many European countries (Mauri et al. 2017). The high degree of genetic diversity and phenotypic plasticity is likely the main reason underlying its broad success; nevertheless until the 1960’s the provenance of the parent seed sources was completely neglected (Bastien et al. 2013). Many national or local experiments have been established using independent seed collections from the native range, mainly according to local climate and expert knowledge.
The first attempt to reorder on a scientific basis the Douglas-fir introduction in Europe was provided between 1965 and 1970 by the International Union of Forest Research Organisations (IUFRO), setting up a broad experimental network of trials in Europe. Adaptive and growth traits of 182 provenances of Douglas-fir (both coastal and interior) from the whole native range (Fig, 1) were tested under a systematic scheme (Eilmann et al. 2013; Isaac-Renton et al. 2014). Provenance trials proved that, under favourable climatic conditions, coastal provenances of Douglas-fir outperformed the interior ones even in continental Europe (Boiffin et al. 2017) and grew faster than native conifers, e.g. Scots pine (Pinus sylvestris L.) or European larch (Larix decidua Miller). Unfortunately, the connection between IUFRO experimental trials (and seed sources) and the older plantations was never considered.
In recent years, the interest in Douglas-fir has risen in Europe mainly due to its ability to cope with chronic drought, making this tree species a valuable option for European forest managers and timber production. Also, Douglas-fir has been sometimes considered a feasible alternative to Norway spruce plantations for pulp production in Central Europe, which recently suffered from drought and pest attacks (Vitali et al. 2017c). Given the assumption that “the geographic origin of tree species is a fair proxy of its genetic diversity” (Eilmann et al. 2013), several studies have been carried out in Europe by means of DNA microsatellite markers to trace back the Central European Douglas-fir stands of unknown origin to their native sources (Van Loo et al. 2019). In many studies several old-growth stands were sampled and compared with IUFRO’s provenances using nuclear SSR (Hintsteiner et al. 2018) as well as with samples collected from the native range. As a result, the coastal variety of Douglas-fir was often the most present, except for a few individuals a posteriori assigned to the interior variety.
Italy is the southernmost European country where Douglas-fir was experimented since the beginning of the XX century (Ciancio et al. 1982) with two IUFRO trials and more than 50 inventoried stands spread along a latitudinal gradient (Castaldi et al. 2017; Marchi and Cocozza 2021). While many European countries studied the old-growth stands of this species (i.e. France, Austria, Germany), this information is completely missing in Italy. In this context, the aim of this project is to fill this gap by means of a combination of molecular markers, spatial sampling algorithms and laser scanning techniques. The Italian old-growth stands will be traced back and the growth and the intraspecific competitive performances of this species will be investigated also under Mediterranean conditions.
An accurate genotype-environment performance assessment is currently missing in many countries including Italy (Marchi and Cocozza 2021). The CONIFIR project aims to assess genotype-level growth and competitive ability of Douglas-fir trees in the Italian stands, and to evaluate their intra-specific diversity and adaptive potential to projected climate change scenarios, as well as to assess their suitability as seed sources for reforestation activities under different climatic regimes. The general goal is to develop a species-tailored research framework by combining geographic, climatic, and molecular data, stand structure, long-term growth trends and predictive models (i.e. Universal Response Functions) for the species in Italy. The final outputs of the CONIFIR project will fill the gap between Italy and many European countries.
To characterise Douglas-fir genotypes currently established in Italy and assess their performance over time and space, a cross-disciplinary approach will be applied and ecological models will be developed. Nuclear SSR markers will be used to untangle the genetic structure of old-growth stands established since the 1850s and to trace them back to their native populations. High-quality data from a precision forestry approach using terrestrial laser scanning technologies will allow us to assess the growth performance and the within-stand competition dynamics of genotypes, as well as their performances in different ecological environments. Combining this tree-level information will allow us to improve our knowledge on the plasticity of the species across Italy and will provide the input data for a robust spatial modelling aimed at predicting the most likely scenarios for Douglas-fir in Italy after the anticipated climate change.
Genetic diversity or intraspecific biodiversity plays a key role for both the long-term survival of a species and the stability and resilience of forest ecosystems, thus affecting their management as well. The current geographic distribution of Douglas-fir stands in Italy will be initially drawn from the most recent National Inventory data (INFC2005 or INFC2015 if already released) which also include tree-level information, though their exact position on the ground is not available (Borghetti & Chirici 2016). According to the species presence (approximately 70 stands throughout Italy), about 35 Douglas-fir stands will be selected as candidate for sampling, taking care of covering the largest variability of climatic conditions as inferred from the most reliable climatic datasets (Marchi et al. 2021, Wang et al. 2016). In each stand, 24 adult individuals will be selected following a spatially-unbiased sampling design, permanently marked and georeferenced to ensure their future availability. From each individual, leaf samples will be collected for DNA extraction; however, due to the remarkable height of many individuals (up to 40 m), samples will be taken from the cambium whenever needed and appropriate technical adjustments will be applied to ensure high-quality sampled material. At the same time, samples for DNA analysis will be collected from a Douglas-fir common garden experiment located in Austria (a collaboration is currently ongoing with Dr. Marcela van Loo, Institute of Silviculture, BOKU University, Vienna), which includes adult trees from 39 populations of known origin spread over the species’ native range in US and Canada. Overall, approx. 1500 Douglas-fir trees will be sampled for genetic analysis in this phase.
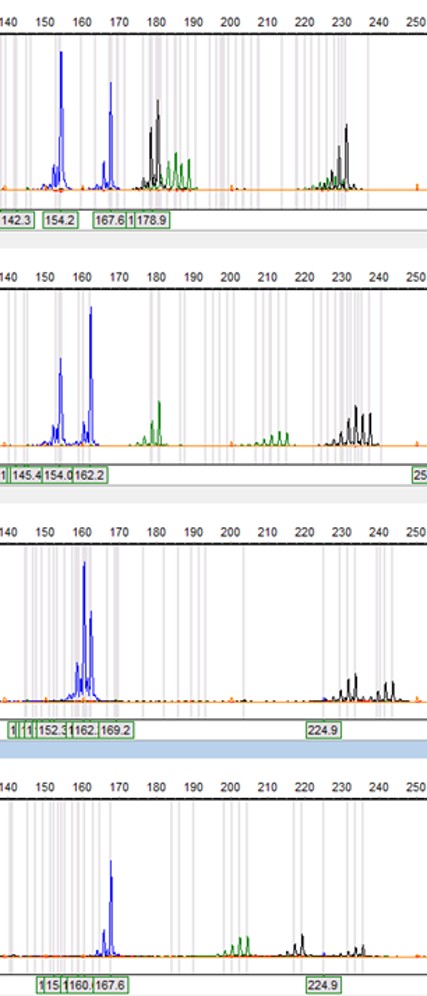 Genotyping of Italian Douglas fir stands. Some 15 polymorphic nuSSRs have been investigated to characterize the genetic makeup of the Italian Douglas fir stands. Here, the electropherograms of 3 out of 15 nuSSRs are shown (blue: PmOSU_2C2; green: PmOSU_2D6; black: PmOSU_1C3) in four trees (top to bottom diagrams). Their relative profiles allowed the discrimination of heterozygote individuals from homozygotes when doubled peaks were detected for the same marker, as shown in the two uppermost PmOSU_2C2 profiles (blue peaks) (courtesy of: P. Iovieno, CNR-IBBR, Italy).
Genotyping of Italian Douglas fir stands. Some 15 polymorphic nuSSRs have been investigated to characterize the genetic makeup of the Italian Douglas fir stands. Here, the electropherograms of 3 out of 15 nuSSRs are shown (blue: PmOSU_2C2; green: PmOSU_2D6; black: PmOSU_1C3) in four trees (top to bottom diagrams). Their relative profiles allowed the discrimination of heterozygote individuals from homozygotes when doubled peaks were detected for the same marker, as shown in the two uppermost PmOSU_2C2 profiles (blue peaks) (courtesy of: P. Iovieno, CNR-IBBR, Italy).
The samples collected from both the Italian stands and the Austrian common garden experiment will be genotyped using 15 highly-polymorphic, unlinked nuclear SSR (Hintsteiner et al. 2018), which will be screened by four multiplex PCR combinations (van Loo et al. 2015). The genetic variation of Douglas-fir populations in Italy will be assessed by computing the main population genetic parameters. Multivariate analysis based on genetic distances will allow the detection of possible clusters of populations/individuals within populations. The genotypes from stands of unknown origin will be assigned to their native provenances by comparing their genetic profile with that of individuals of known origin maintained at the Austrian common garden experiment. This will allow us to trace back each Italian population to any of the 12 genetic clusters recently identified by Hintsteiner et al. (2018), using an adequate number of reference native populations. A Bayesian clustering approach will be applied to assign each population/individual to either Douglas-fir variety (coastal/rocky mountains) and to detect cases of artificial intervarietal admixture in the Italian stands. Further analyses will be carried out within each variety at each of the four hierarchical levels identified by Hintsteiner et al. (2018) to assign each of the Italian stands to one of the reference North American populations within each of the 12 genetic clusters. Additional assignments will be performed at individual level to identify possible cluster-admixed stands. Within-population genetic diversity indices will allow to describe the current genetic structure of italian stands and to devise several best practices for genetic-based management and regeneration of the species.
An optimal sampling strategy is required to fully capture the extant genetic diversity at both inter- and intra-population level, as well as to optimise the choice of populations/mother trees to be marked as seed sources, thereby maximising the genetic diversity and the potential adaptability of the next-generation trees. Survey sampling in forest field inventory traditionally considers classic sampling methods such as simple random sampling, systematic sampling and spatial stratification. More recently, several spatial sampling methods have been proposed in the literature (Benedetti et al. 2017) which allow the selection of the units incorporating their geographical positions at the design phase, leading to more accurate estimates of the population parameters. Such strategies have been tested on real populations (Grafström et al., 2017) but no distinctions have been made among natural and artificial forest landscapes. Indeed, natural populations follow expected structures and dynamics and both classical and spatial sampling techniques are widely and efficiently used in capturing the genetic diversity. As for artificial populations like Douglas-fir plantations, no spatial genetic structure is expected to exist in the current stands as a result of the nursery material mixing during harvesting, breeding, transporting and planting the seeds. However, depending on the original propagation material, the planting scheme, and the long-time intraspecific competition among different genotypes in contrasting habitats, the current stand genetic make-up could deeply differ among Douglas-fir stands. This leads to carefully considering the sampling techniques for tree selection with the aim of reducing the sampling costs. After the collection of the information on the whole stands, simulation studies may be conducted firstly to understand the real possibility and result quality in applying existing sampling methods, and then to develop a specific strategy, which mixes random and ad-hoc selection of trees, e.g. spatially balanced techniques mixed with indirect or adaptive ones. Simulation results based on sampled trees genotyped for nuSSR will allow to find evidence that recommend the use of a feasible and less costly sampling design suitable for such artificial stands as well as recommendations on the best practices for optimising the selection of seed sources. In addition, an in-depth study will be conducted to understand the optimal sample size to capture the within-stand competition dynamics, in order to reduce costs in the survey implementation.
Tree architecture and growth rate are strongly influenced by the competition dynamics among neighbouring trees which impacts individual growth rate and mortality rate. New developments in terrestrial laser scanning (TLS) provide unprecedented three-dimensional in situ information of trees and forests. TLS is a ground-based laser scanning technology which can provide detailed three-dimensional information to precisely depict forest structure in a non-destructive, objective, and reproducible manner. Compared to traditional methods, the use of TLS allows the collection of massive amounts of point cloud data for quantifying three-dimensional habitat properties at increasing spatial and temporal scales. The potential of TLS for forest monitoring was first demonstrated in published literature in the early 2000s. Applications initially focused on measuring traditional structural metrics commonly used in forestry such as height and diameter at breast height (dbh), but eventually evolved to whole-tree volumetric assessment to improve estimates of aboveground biomass. Current applications also include individual modelling of tree architecture and habitat assessment.
While the use of TLS for measuring tree horizontal and vertical forest stand attributes has been widely explored, additional outputs can also include three-dimensional reconstructions of individual tree structure (Puletti et al. 2019) or vegetation profiles (Ashcroft et al. 2014). In addition, TLS holds strong potential in retrieving stand competition dynamics due to its ability to precisely measure both the horizontal and vertical tree crown structure and arrangement from the 3D point cloud. TLS data will be acquired using a FARO Focus 3D × 130 (FARO Technologies Inc., Lake Mary, FL, USA). The instrument uses a phase-shift-based technology with a maximum range of 120 m and acquires data with an azimuth scan angle of 360°. It collects the x, y, and z coordinates and the intensity of the laser returns with a scan ranging noise of ±1 mm (FARO User Manual, 2018). A complete description of the instrument can be found in Giannetti et al. (2018).
The specific objective is to reconstruct the single-tree architecture by TLS point cloud data and evaluate its relation with the competitive ability of genotyped trees. The reconstruction procedure had two main components: (a) manual tree extraction from the registered point cloud, (b) 3D reconstruction of individual tree point clouds. In order to achieve this task, 3 Douglas-fir stands across three different ecological regions (according to climatic analysis) will be selected for extended investigations among those previously characterised by genetic analysis. In each of the 3 stands, an area including at least 24 trees previously genotyped (see above) will be laser-scanned and 3D reconstructed. The following attributes will be considered and measured: dbh, tree total height, stem length from the ground to the first big branch (d > 20 cm), crown features (volume, deepness, spread), slenderness coefficient, dead wood presence (standing and lying dead trees, coarse woody debris, snags, stumps, etc.) and best assortment achievable (sawn timber, veneer, board, etc.). Winkelmass, dbh-dominance, Hegyi index and Hegyi index-modified will be considered as tree-level competition indices, among others.
Old-growth (>70 years old) Douglas-fir populations established across Europe represent one of the largest transfers of tree genetic material outside its natural range (Boiffin et al. 2017). When tree species are transferred, genotypes are subjected to non-analogue environmental conditions, which are sometimes very different from those of the native range. The larger the shift is, the more interesting and uncertain the tree responses are. In a climate change scenario, the intentional movement of tree species can provide important data on how tree species can react to different environments. While some central-european countries worked on this topic, few research has been performed in Southern Europe; for this reason models including genetic data are required in the Mediterranean area.
The ecological characterization of Douglas-fir stands in Italy will be used in CONIFIR to define the “multivariate ecological distance” in between them and their parent populations in the native range. This will allow us to infer the ecological plasticity of each genotype sampled based on its geographic distribution across different climatic regimes. To achieve this a downscaling tool named ClimateDT (freely available at https://ibbr.cnr.it//climate-dt/) will be used. This system represents an improvement of ClimateEU (Marchi et al. 2020) and other softwares for other parts of the globe such as ClimateNA (Wang et al, 2016, Liu et al. 2018). The core of ClimateDT is a dynamic downscaling approach where local lapse rates are used to take into account the influence of altitude after a bilinear interpolation. As a common feature, ClimateDT is based on CRU-TS v.4.05 dataset (Harris et al. 2020), one of the most well referenced and used in ecological studies, often employed to reconstruct historical climate worldwide. The dataset has a monthly resolution spanning from 1901 to 2019 but its original spatial resolution (0.5°, ~50km at the equator) is too coarse for regional studies. For this reason downscaling processes are necessary for local studies. In addition to historic climate, ClimateDT also offers more than 80 climatic variables and future scenarios with monthly resolution over the period 1901-2098. The future climates are derived from the global UKCP18 dataset (~60km at the equator), which comprises 28 different GCMs (named “variants”) for 2 RCP trajectories from CMIP5 and 6 GCMS for 4 ssp scenarios from CMIP6.
The complete climatic dataset provided by ClimateDT (1901-2098) will be used to study the effect of climate in the future and to predict possible future scenarios. In combination with climatic analysis, a Universal Response Function (URF) will be developed for Douglas-fir across Italy. According to Wang et al. (2010) these models are a combination of individual response functions and individual transfer functions. Individual response functions are used to model a trait measured for a specific tree from a specific population in a single test site. The trait is modelled as a function of local climate (i.e. the climate at the planting site or common garden experiment) and the final output is the expected performance at that specific site. In contrast, individual transfer functions have been developed to study the expected increase or decrease in growth when a species is planted along an ecological gradient. Yet, simple linear regression models used for fitting have been often questioned due to the need of including in the model the genetic component and other trials features more properly (Benito Garzón et al. 2019, Marchi et al. 2022).
In CONIFIR, URF models will be developed by combining the trait measurements with climate shifts and genetic information. A mixed-effect quadratic URF will be built for Douglas-fir in Italy consistently with previous studies using climatic variables of test sites and provenances as independent variables. Both environmental effects of local climate and genetic effects (among-population variation) of the climate in the native range on phenotypic performance will be evaluated and integrated into a single model. However, it is also known that among-population variation can be considerably affected by other factors such as gene flow, demographic history, and colonisation events, resulting in spatial patterns that are independent of climate. These spatial genetic effects are somehow absent in Italy due to the artificial origin of Douglas-fir stands, thus literature data will be used to derive suitable information from the native range. In the CONIFIR project the novel information collected by the different tasks (i.e. genetic+climate+growth) will be combined to calibrate and validate a URF aimed at obtaining spatial predictions of habitat suitability and growth performances for Douglas-fir across Italy. The URF approach which models tree growth as a function of both environmental (planting site) and genetic (seed source) effects will be applied, allowing the growth of any provenance to be predicted at any location (Wang et al. 2010).