
CONIFIR Project
Genetic origin and structural setting of Douglas-fir planted forests in Italy for their management, conservation and valorization
MUR-PRIN 2022 Project - Prot. 20225WRLS3

MUR-PRIN 2022 Project - Prot. 20225WRLS3
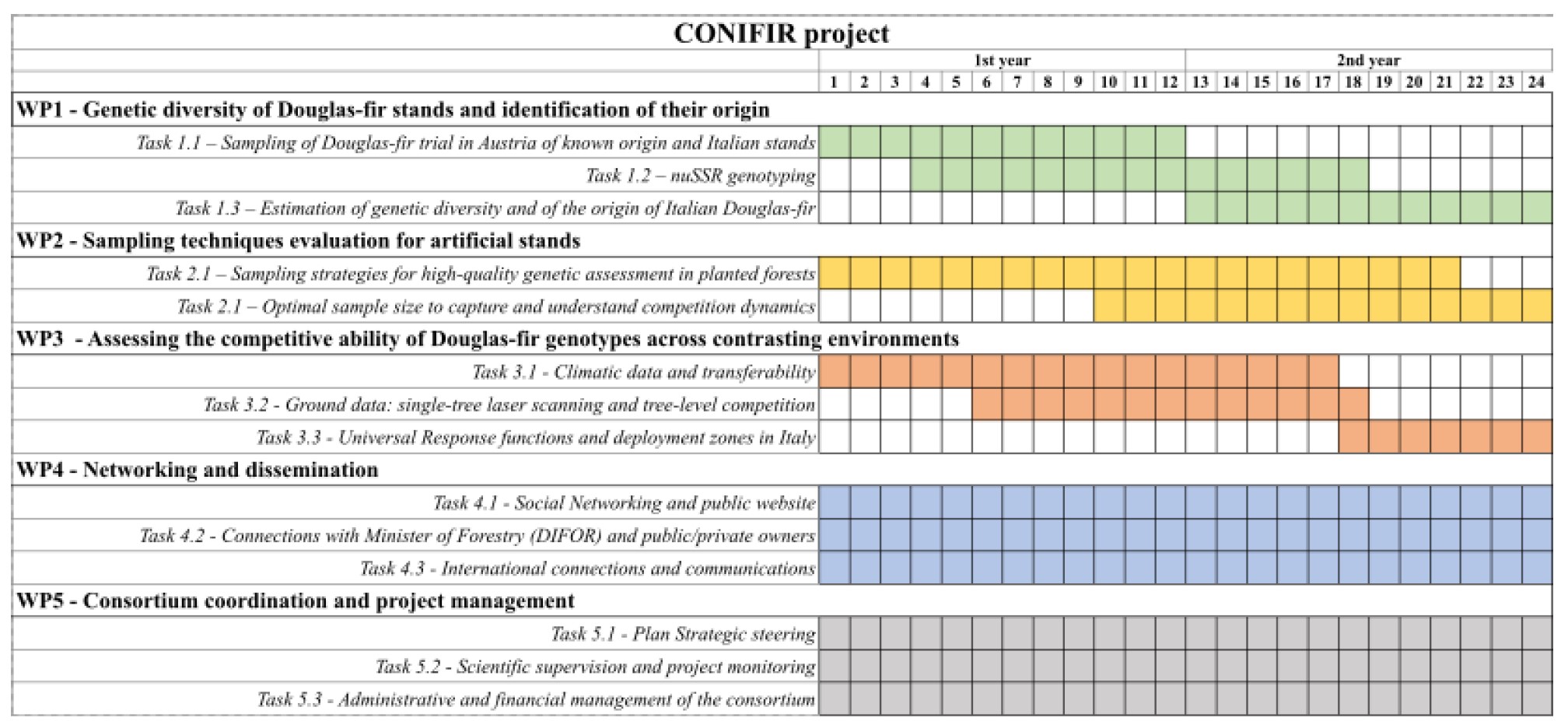
The Gantt chart of the project.
(Leader: Paolo Iovieno - Months: m1-m18)
The WP1 will characterise the genetic make-up of Italian Douglas-fir populations of unknown origin and will estimate the genetic variability of them. Additional work will be done on the 3 stands which will be intensively investigated for stand structure characteristics and competition dynamics.
The Austrian field trial at BOKU University, which hosts Douglas-fir trees of known origin (see above), will be sampled for DNA analysis and used as reference to trace back the Italian Douglas-fir stands of unknown origin to their parent native populations. At the same time, the most representative Italian stands (with Douglas-fir basal area > 65% according to INFC2005) will be sampled: 40 stands are estimated to match the above criteria. In each stand, 24 adult individuals will be selected following a spatially-unbiased sampling design.
Twenty-four individuals will be genotyped in each selected stand using 13 nuclear SSR markers, retrieved by literature for their high polymorphism, together with the genomic DNAs of the selected reference native populations. Several additional individuals surrounding the 24 trees sampled in Task 1.1 will be screened using the same SSR markers to investigate the within-stand competition dynamics among different genotypes/provenances in the 3 stands selected for extended sampling (see above).
The dataset obtained from genotyping in Task 1.2 will be analysed by advanced statistical techniques in order to trace back Italian stands and to evaluate the different sampling schemes proposed in Task 2.1. A Bayesian approach recently implemented with Central European Douglas-fir populations will be used.
(Leader: Maria Michela Dickson - Months: m1-m24]
The WP2 will work on a retrospective analysis aimed to capture the real extent of the genetic diversity between and within populations, by selecting a sample that can noticeably reduce costs. Moreover, since competition inside a forest stand constitutes a non-negligible issue, as faced out in WP3, a relevant problem is the search for an optimal sample size that can allow the phenomenon to be accurately studied without resorting to the complete visiting of all the units in the population. Both issues can lead to indications and best practices in studying artificial stands.
A complete Monte Carlo simulation study will be conducted using several sampling techniques, both traditional and spatially balanced. The possibility to implement a new, more efficient sampling strategy for artificial stands will be explored and developed, both from the statistical and practical point of view. The quality of the results will be evaluated in terms of performances in total and mean estimation of the studied characteristics by comparing simulation results to the observed values collected in WP1.
The dynamics of competition at tree-level could be captured considering samples of units designed to mirror the real situation. Such samples need to be defined using appropriate sampling designs, allocations and sizes. Starting from the collected data, Monte Carlo simulations will be implemented in order to define optimal features of the sampling strategy.
(Leader: Maurizio Marchi - Months: m6-m24]
The WP3 will develop trait-genetic diversity models to meet durable adaptation and production of Douglas-fir stands in Italy.
Downscaled climate time series from ClimateDT will be generated for the native range and the Italian planting sites in order to evaluate the climatic shift that Douglas fir provenances have experienced in Italy. The climate of origin in the native range will be reconstructed as long-term normal climate (i.e. 1901-2020) while the average climate for the whole life-span of the stand will be used for planting sites and adjusted site by site.
The objective of this task is to create a database of approximately 70 laser-scanned trees of known genotype (see Task 1.2). Single tree-architecture models will be generated using TLS, together with other tree-architectural attributes like, e.g. dbh, total height slenderness ratio, crown features, etc. Competition indices will be computed in order to quantify the tree-level competition among neighbouring trees.
Response functions will be created to provide measures of short-term acclimation and specific responses to climatic drivers. Reaction norms will be based on environmental and genetic data to identify major genetic (G) and environmental (E) determinants, as well as the part of phenotypic plasticity under genetic control (GxE).
(Leader: Gabriele Bucci - Months: m1-m24]
The project outcomes will be properly packaged to reach, engage and address the needs of the final users and widely shared with relevant stakeholders involved in the optimal management and exploitation of forest tree species.
Timely regular blog posts and annual digests will be released and widely distributed. The CONIFIR public website will be implemented as a dynamic communication tool, including: i) a showcase of the project objectives, work plan, ongoing activities and results; ii) the publication of regular project and sector-related news; iii) a portal for the tools, models, protocols, guidelines, best practices and surveys, and other deliverables; and iv) links to other EU or international forest initiatives, projects and stakeholder associations.
A list of key contacts will be assembled to spread the information to a wide audience in Italy, including stakeholders, politicians and Ministers. Scientific, forestry and industrial associations, scientific societies, policy makers, journalists, national agencies in the forest sector, etc. will be informed about the project achievements through key communications and dissemination channels.
CONIFIR will develop tools and facilities to link Italian, European, Canadian and American researchers aimed at evaluating the performance of Douglas-fir in different ecological ranges. The connection of the project dataset with national and international research infrastructures (RI) will help realise this task.